BQM 329
The noseki continues with white using a tesuji to escape with the  group to the center. White has a weak group, but it seems to me that
group to the center. White has a weak group, but it seems to me that 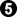 is a slow move.
is a slow move.
Dave: Personally I think that
is natural. To me there is a question whether
tries too hard (see comment 2 diagrams down the page). I would be inclined to simply play a and capture the White stones on a large scale or attack them if they try to escape.
Dave: Am I blind here? What happens if White plays 2 at 5? Black can not push through and cut, White has too many liberties, no? If Black plays 2, White 4, Black 7, and White 6 is this worse than the dumpling?
Bill: Good morning, Dave. You are correct about
. That's more evidence that there is a Black stone at
. If nobody objects, shall we put one there in the original diagrams? I do not think that it affects the BQM itself.
tderz: Bill, wouldn't that rather be two separate threads?
- one var with a black stone at

- one var without
 =
= 
Bill: The BQM was actually about what happened if Black captured the stone on the left instead of making a bamboo joint. I think that that question is the same, regardless of whether there is a stone at
or not. However, there is also a lot of discussion already about the play in other variations where it matters a great deal. Maybe we could make that discussion a subpage, or a new BQM?
Dave: I think we should at least keep the original diagram as is. Basically the cut looks wrong but it also looks like just the kind of annoying play that stronger players occasionally pull on us hapless beginners at the worst possible moment. :-)
Bill: Since Imagist is relying on memory (See below.), it seems to me that this variation is enough to show the problem with  in the original diagram and the adequacy of
in the original diagram and the adequacy of 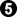 in the second diagram. White's dumpling is clearly inefficient.
in the second diagram. White's dumpling is clearly inefficient.
Dave: I think it will depend on the surrounding situation. However, since Black chose to block on the top against the original 3-3 invasion, the top is presumably important. While White's dumpling is inefficient, if we add the 4 - 6 exchange, is it so clear that the result is bad for White?
Honte & shape destroying tesujis
Go literature that I have read seems to indicate that black 15, (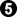 in the diagram to the left) is a bad move.
in the diagram to the left) is a bad move.
The reason seems to be the tesuji with  -
- . But, my question is, can white really seal black in? And even if she can seal black in, can she win the resulting capturing race?
. But, my question is, can white really seal black in? And even if she can seal black in, can she win the resulting capturing race?
Dave: But in this case isn't
sente after 1 through 9 so White can both live in the corner and on the outside for a very good result from a 3-3 invasion?
pwaldron: Good catch Dave. Another fast thought: how about playing  instead of the hanging connection at a? It seems that it reduces the threat of White activating the outside stone, giving Black time to capture with
instead of the hanging connection at a? It seems that it reduces the threat of White activating the outside stone, giving Black time to capture with 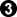 (haven't checked this for sure, though). First impression is that Black ends up a little thicker on the outside, with no improvement for White over the standard line.
(haven't checked this for sure, though). First impression is that Black ends up a little thicker on the outside, with no improvement for White over the standard line.
Dave: Agreed but...
Dave: What if this annoying so-and-so goes ahead and plays  anyway? If Black captures in the corner, White can force Black along the top edge with
anyway? If Black captures in the corner, White can force Black along the top edge with  .
.
tderz: Quite sure that it would be a very bad attempt to crawl on the 2nd line (a-b-...?) ,
however I would like to know in how many moves could one capture the corner (m),
resp. how many liberties can allow White to Black's crawling group before hane-ing?
Dave: If  and
and  ,
,  is not going to work against
is not going to work against  since the White enough liberties to capture the 3 Black stones if Black tries to push through and cut.
since the White enough liberties to capture the 3 Black stones if Black tries to push through and cut.
Charles  is surely wrong. Even if you wanted to play this way as White, you'd play
is surely wrong. Even if you wanted to play this way as White, you'd play  first and get the answer.
first and get the answer.
Second point: the shutting in isn't absolute: Black can get out with an odd play.
tderz: As Charles points out above,
it is good style to decide late which cutting point (d, e?) to take.
This way, one could interprete a here ( in uppermost diagram) as yosumiru (have-a-look-how-you-will-react-) move.
in uppermost diagram) as yosumiru (have-a-look-how-you-will-react-) move.
- d: if W plays d first, Black is free to decide how to deal with cutting point c after W hane-d at a.
- e: if W first cuts (ataris) at e,
then after the exchange connect d, atari f - g - capture h - corner securing i, W has exchanged the corner for a low side ponuki.
There is still the common corner aji (Wk - Bl) left, which enables W to push up at m and Black must be very careful about aji around o and p.
tderz: delaying the choice of cutting points enables W to cut at  .
.
This is espec. nice if there is a (high) (handicap) black stone at  , as this will become devaluated.
, as this will become devaluated.
tderz:  at x.
at x.
for the territorial players  is an option.
is an option.
Because  delayed the choice between d,e in an above diagram,
delayed the choice between d,e in an above diagram,
White can now get a stone , while not losing the corner.
(exchange  -p follows)
-p follows)
tderz: Black plans to go territorial by protecting both cutting points e,d simulateously with  .
.
....
tderz: ...this is true , and indeed , after B collected 6 liberties on the 2nd line (!) he can go for the corner with  +
+  .
.
However, in most cases the influence gained by White will outweigh the loss of the corner.
Imagine a black handicap stone at  ! (would be totally wasted)
! (would be totally wasted)
tderz: Given the double-hane  here,
here,
another comment I would like to make is,
that IMO, one does not always have to protect cutting point c in a tight way, i.e. here black d.
- Firstly one knows that Black can live anyway,
- secondly
 might have been played for a position on the lower side.
might have been played for a position on the lower side.
Dave: Presumably if the key to this position were on the lower side, Black would block on the lower side against the original 3-3 invasion. If Black went ahead and blocked as here anyway then played the marked stone in relation to a lower side position, White would not have made any noseki cut but rather simply cut to the right of the marked stone and happily captured it in the normal joseki, no?
tderz:  enables to control
enables to control  with geta g (=
with geta g (= ).
).
IMO, Black is happy about the greedy & hasty cut  +
+  as Black sacrifices the stone for influence.
as Black sacrifices the stone for influence.
The sequence could end with either  or connection h.
or connection h.
tderz: Black can get lower & outside influence by the nice move of  which doesn't give W so many cutting points.
which doesn't give W so many cutting points.
fractic:  looks a bit odd but if we add one black stone it's pretty common.
looks a bit odd but if we add one black stone it's pretty common.
fractic:  at a and 12 at b. Now both Kogo's Joseki dictionary and Tesuji by Davies mention that white needs a stone at the circled point in order to be able to play at c. With that stone they claim that the tesuji combination would ineed succeed and kill black. Personally I can't find out how and allways seem to fall one liberty short for white. Also fuseki.info only has one game with this varation indeed with a white stone at the circled point.
at a and 12 at b. Now both Kogo's Joseki dictionary and Tesuji by Davies mention that white needs a stone at the circled point in order to be able to play at c. With that stone they claim that the tesuji combination would ineed succeed and kill black. Personally I can't find out how and allways seem to fall one liberty short for white. Also fuseki.info only has one game with this varation indeed with a white stone at the circled point.
Imagist: Sorry, I was trying to do this from memory since I didn't have a book handy when I was writing this; the joseki in question is indeed the one shown by frantic? above. I apologize for the confusion.
I still can't find the tesuji or joseki books that I got this from, but I found a similar situation where Kageyama insists that  and
and  seal black in and capture (without any mention of a stone along the side):
seal black in and capture (without any mention of a stone along the side):
Kageyama says that white answers black a with white b.
fractic: White is allready alive in the corner in this diagram.
![[Diagram]](diagrams/23/e512fec931b5c399452588c68555665a.png)
![[Diagram]](diagrams/9/421215bb97da5834e9181ed6138e6a62.png)
![[Diagram]](diagrams/10/702147fe8e6e18fdd2ab58a9fca5b7ac.png)
![[Diagram]](diagrams/32/bd23b1030a0755be449e77675b111651.png)
![[Diagram]](diagrams/31/eedaae1de812cb3445445ffc514b273b.png)
![[Diagram]](diagrams/12/da0f631fc9fc549b7035b2013e079240.png)
![[Diagram]](diagrams/24/911f2f446954745bcfe24cd9c18ae8cb.png)
![[Diagram]](diagrams/24/dabe10a04086a2b9d8d1b485e4028e88.png)
![[Diagram]](diagrams/48/8915437071373ba3c20aa29644b37002.png)
![[Diagram]](diagrams/36/348ebc727ccd2e9093f04a2dbbd165b5.png)
![[Diagram]](diagrams/50/8fbbf4f76bb04ac3e3a2406a77557a47.png)
![[Diagram]](diagrams/52/6507e4451813484b3281fa05d6db2e5a.png)
![[Diagram]](diagrams/19/71932705574653661bb9fb0e1fc32644.png)
![[Diagram]](diagrams/38/a0b0c0cc25655f982e5660a90f29c704.png)
![[Diagram]](diagrams/49/7882a2c1f400ffd5d5352892c58d44c4.png)
![[Diagram]](diagrams/39/ad6d6864a3dc989e8ae43c7513220e6a.png)
![[Diagram]](diagrams/17/9289f0cca1fceb58bfe5c1209363d778.png)
![[Diagram]](diagrams/43/b9d4276d71b20ba72efdb14fc84d69fc.png)
![[Diagram]](diagrams/35/ffea0d490a2496f60ba113ddb4bb54e1.png)
![[Diagram]](diagrams/37/396bd7223aea12d3a601cdbeb3eabcc4.png)
![[Diagram]](diagrams/15/456d0450b0605acb830ee89d57b33b96.png)
![[Diagram]](diagrams/4/ae8a10a82a40e88d5365f2b35463580a.png)
![[Diagram]](diagrams/51/326cfcfa91f1b9838848435f1be83929.png)
![Sensei's Library [Welcome to Sensei's Library!]](images/stone-hello.png)